Throughout this project, I aim to adress three of the long-standing questions in speciation research:
1) What constitutes the genetic architecture of reproductive barriers?
2) To what extent do chromosomal rearrangements contribute to speciation?
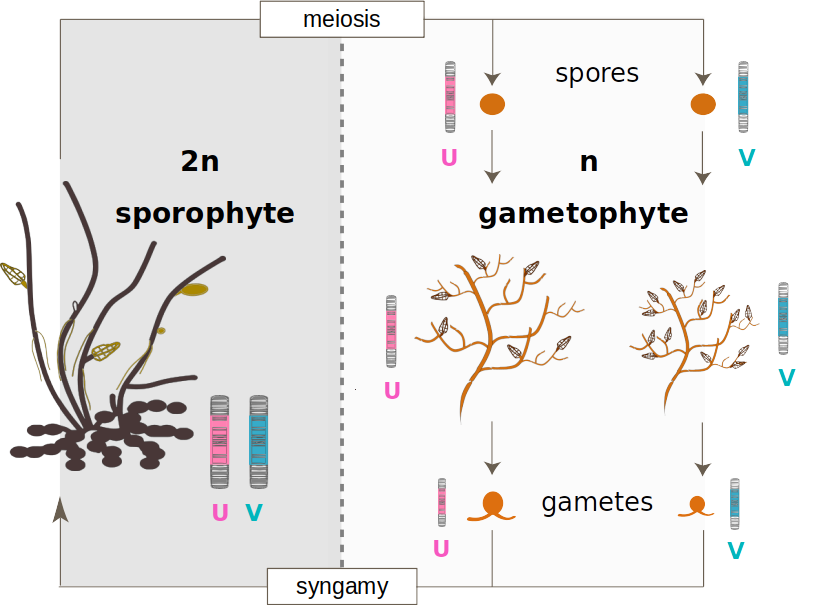
3) What roles do haploid sex chromosomes play in the evolution of reproductive isolation?